GenomicRangeQuery
Test description
A DNA sequence can be represented as a string consisting of the letters A, C, Gand T, which correspond to the types of successive nucleotides in the sequence. Each nucleotide has an impact factor, which is an integer. Nucleotides of types A, C, G and T have impact factors of 1, 2, 3 and 4, respectively. You are going to answer several queries of the form: What is the minimal impact factor of nucleotides contained in a particular part of the given DNA sequence?
The DNA sequence is given as a non-empty string S = S[0]S[1]...S[N-1]consisting of N characters. There are M queries, which are given in non-empty arrays P and Q, each consisting of M integers. The K-th query (0 ≤ K < M) requires you to find the minimal impact factor of nucleotides contained in the DNA sequence between positions P[K] and Q[K] (inclusive).
For example, consider string S = CAGCCTA and arrays P, Q such that:
P[0] = 2 Q[0] = 4 P[1] = 5 Q[1] = 5 P[2] = 0 Q[2] = 6
The answers to these M = 3 queries are as follows:
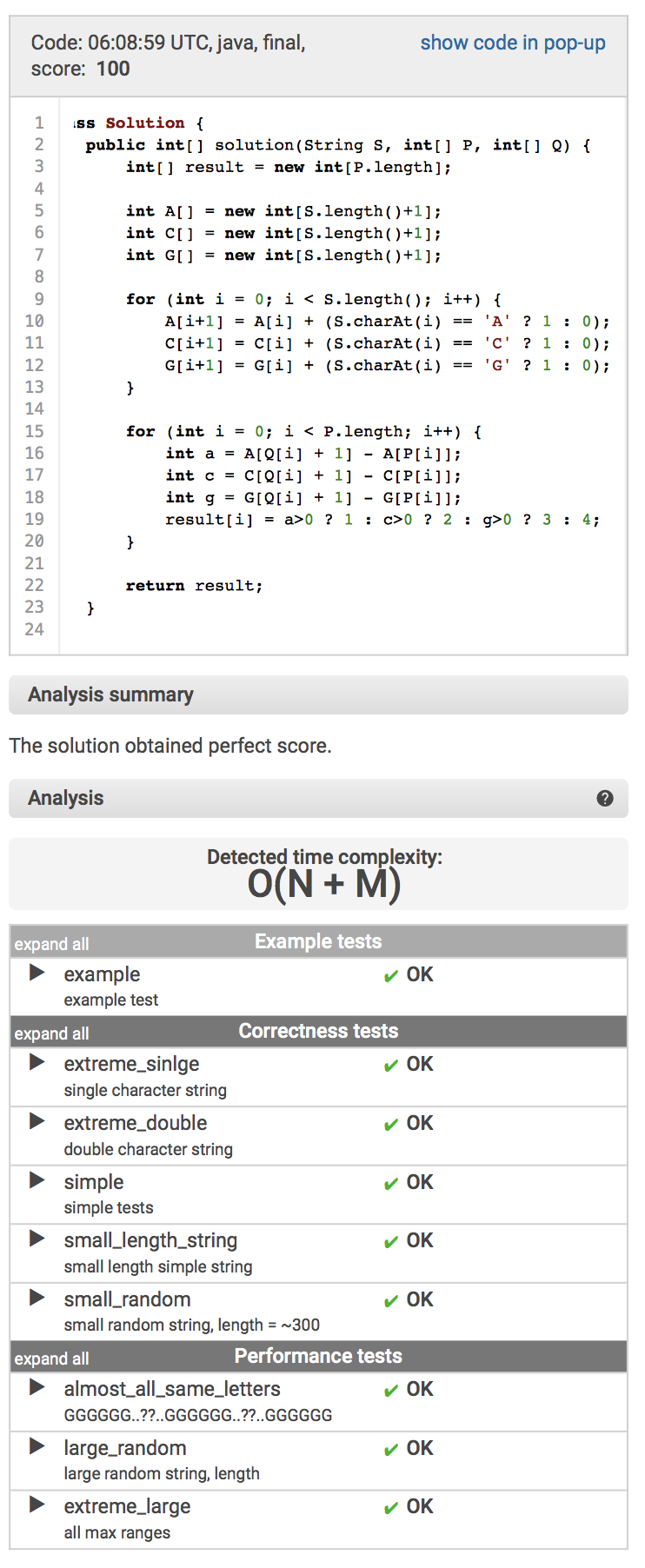
class Solution { public int[] solution(String S, int[] P, int[] Q); }
that, given a non-empty zero-indexed string S consisting of N characters and two non-empty zero-indexed arrays P and Q consisting of M integers, returns an array consisting of M integers specifying the consecutive answers to all queries.
The sequence should be returned as:
P[0] = 2 Q[0] = 4 P[1] = 5 Q[1] = 5 P[2] = 0 Q[2] = 6the function should return the values [2, 4, 1], as explained above.
Assume that:
A DNA sequence can be represented as a string consisting of the letters A, C, Gand T, which correspond to the types of successive nucleotides in the sequence. Each nucleotide has an impact factor, which is an integer. Nucleotides of types A, C, G and T have impact factors of 1, 2, 3 and 4, respectively. You are going to answer several queries of the form: What is the minimal impact factor of nucleotides contained in a particular part of the given DNA sequence?
The DNA sequence is given as a non-empty string S = S[0]S[1]...S[N-1]consisting of N characters. There are M queries, which are given in non-empty arrays P and Q, each consisting of M integers. The K-th query (0 ≤ K < M) requires you to find the minimal impact factor of nucleotides contained in the DNA sequence between positions P[K] and Q[K] (inclusive).
For example, consider string S = CAGCCTA and arrays P, Q such that:
P[0] = 2 Q[0] = 4 P[1] = 5 Q[1] = 5 P[2] = 0 Q[2] = 6
The answers to these M = 3 queries are as follows:
- The part of the DNA between positions 2 and 4 contains nucleotides G and C (twice), whose impact factors are 3 and 2 respectively, so the answer is 2.
- The part between positions 5 and 5 contains a single nucleotide T, whose impact factor is 4, so the answer is 4.
- The part between positions 0 and 6 (the whole string) contains all nucleotides, in particular nucleotide A whose impact factor is 1, so the answer is 1.
class Solution { public int[] solution(String S, int[] P, int[] Q); }
that, given a non-empty zero-indexed string S consisting of N characters and two non-empty zero-indexed arrays P and Q consisting of M integers, returns an array consisting of M integers specifying the consecutive answers to all queries.
The sequence should be returned as:
- a Results structure (in C), or
- a vector of integers (in C++), or
- a Results record (in Pascal), or
- an array of integers (in any other programming language).
P[0] = 2 Q[0] = 4 P[1] = 5 Q[1] = 5 P[2] = 0 Q[2] = 6the function should return the values [2, 4, 1], as explained above.
Assume that:
- N is an integer within the range [1..100,000];
- M is an integer within the range [1..50,000];
- each element of arrays P, Q is an integer within the range [0..N − 1];
- P[K] ≤ Q[K], where 0 ≤ K < M;
- string S consists only of upper-case English letters A, C, G, T.
- expected worst-case time complexity is O(N+M);
- expected worst-case space complexity is O(N), beyond input storage (not counting the storage required for input arguments).
Answer